CRTC I2M
Contents
Introduction
This document contains instructions for downloading and using the crtc_i2m software suite developed by the CRTC lab at Old Dominion University.
The suite contains three software components:
- Tessellate3D (PODM 3D) (3D tessellation software)
- Tessellate2D (Triangle) (2D tessellation software)
- Convert Image (image format conversion software)
- PartonsToNRRD (creation of image out of Partons data software)
Problem Domains
The image-to-mesh conversion software suite that the CRTC has developed can be used in many different problem domains such as Medical Image Computing and Computational Nuclear Femtography. Specific instructions on how to use the software are described below.
Medical Image Computing
Tessellate3D
In Medical Image Computing, Tessellate3D can be used to generate meshes from multi-tissue segmented images.
Examples
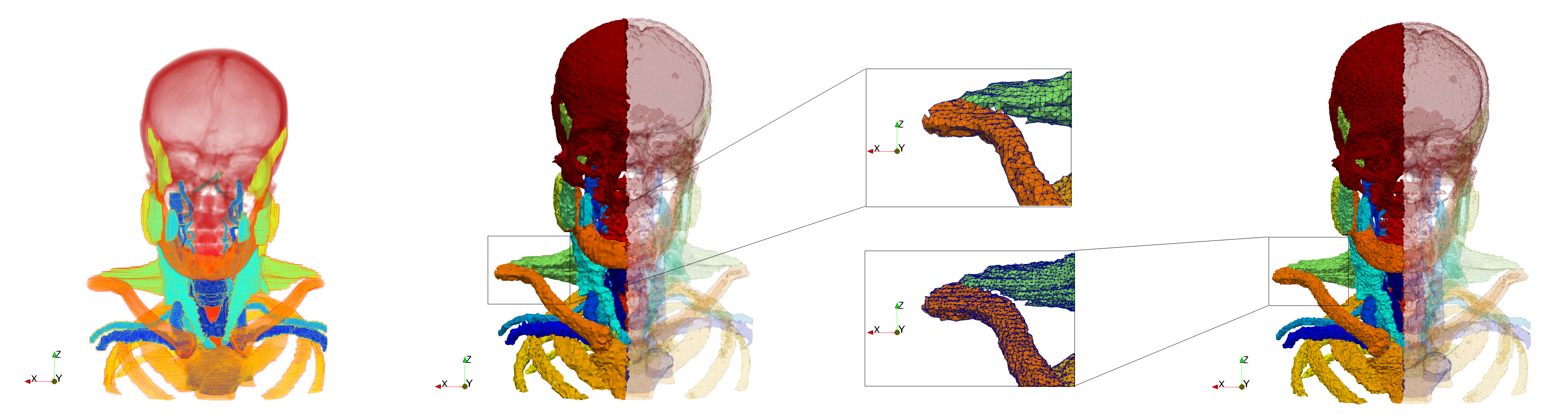
Input image and clipped generated meshes
- Input (left figure): Dimensions (255x255x229) with spacing (0.976562x0.976562x1.40002)
- Uniform with Delta = default (middle figure): 205,510 tetrahedra
- Uniform with Delta = 1.5 (right figure): 767,393 tetrahedra
Uniform with Delta = default (middle figure):
docker run -v $(pwd):/data/ crtc_i2m tessellate3d --input ./Medical_Imaging_Data/3D/Head-Neck.mha --output ./Head-Neck,d=2.49023.vtk
Uniform with Delta = 1.5 (right figure):
docker run -v $(pwd):/data/ crtc_i2m tessellate3d --input ./Medical_Imaging_Data/3D/Head-Neck.mha --delta 1.5 -output ./Head-Neck,d=1.5.vtk
More Detailed 3D examples of Medical Image Computing can be found here.
Parameters
In this domain, some of the parameters of Tessellate3D take on different meanings. For the full range and descriptions of parameters please see the software documentation section of Tessellate3D.
-i, --input [filename] (required)
Input data. It could be an ASCII/Binary NRRD file, an image saved in .vtk format or any other format supported by the ITK library. For more information regarding the file formats supported by the ITK library, see here.
-o, --output [filename] (optional)
The filename of the output mesh saved in VTK format. (Default: input filename + parameters.vtk).
-c, --plc [filename] (optional)
If given, the surface of the produced mesh will be saved into filename in the VTK format.
-t, --threads [unsigned integer] (optional)
Sets the number of threads to be utilized.
(Default: 1).
-d, --delta [unsigned real] (optional)
Controls the density of surface approximation. Smaller values will lead to denser approximation close to the surface (and often to a more accurate surface representation), but will also lead to greater mesh size. The same delta value is used for every tissue of interest. A smaller delta value should be used if at least one of the tissues of interest is not recovered after the Meshing Procedure.
The maximum suggested value for delta is 1 / 5 of the minimum-physical-size of the input image, where minimum-physical-size = min(spacing * size).
(Default: 1 / 100 * minimum-physical-size).
-g, --volume-grading (optional)
Enables the grading of the volume of the mesh. By default, the value of delta controls both the surface approximation and the size of the elements. Using this flag the value of delta will control only the surface approximation resulting in elements of higher volume inside the domain.
--include-labels [unsigned char list] (optional)
Labels that will be included in the mesh. If not specified, all labels will be included in the mesh. Cannot be used together with --exclude-labels.
--exclude-labels [unsigned char list] (optional)
Labels that will be excluded from the mesh. Keeps all labels in the mesh except the ones specified. If not specified, all labels will be included in the mesh. Cannot be used together with --include-labels.
Tessellate2D
Examples
Input image and generated meshes
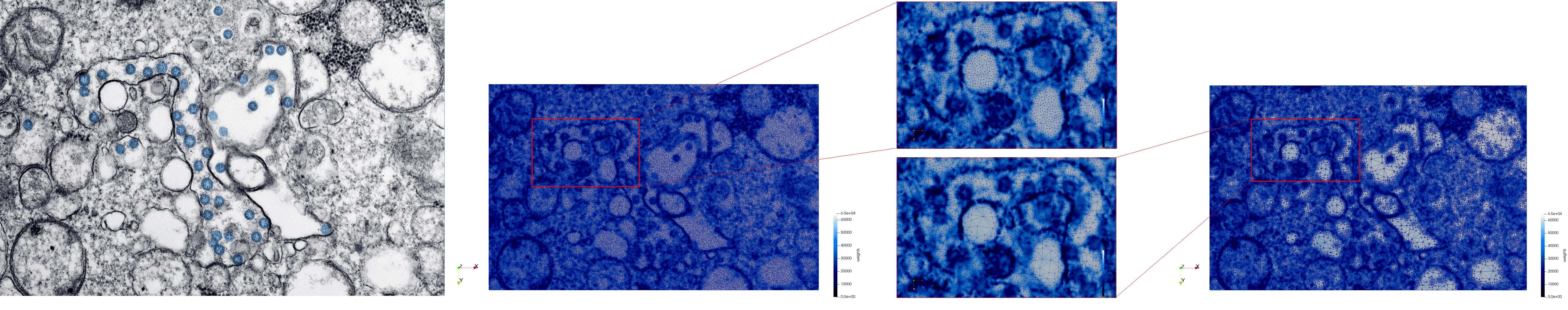
- Input (left figure) : Dimensions (3,000x2,000) with spacing (1x1)
- Uniform with Min-Edge = 15 (middle figure) : 82,981 triangles
- Adaptive with Min-Edge = 15 (right figure) : 67,920 triangles
Uniform with Min-Edge = 15 (middle figure):
docker run -v $(pwd):/data/ crtc_i2m tessellate2d --input ./Medical_Imaging_Data/2D/COVID-19-23354.tif --uniform --min-edge 15 --output ./COVID-19-23354,uniform,e=15.vtk
Adaptive with Min-Edge = 15 (right figure):
docker run -v $(pwd):/data/ crtc_i2m tessellate2d --input ./Medical_Imaging_Data/2D/COVID-19-23354.tif --min-edge 15 --output ./COVID-19-23354,w=0.1,e=15.vtk
Meshes generated based on the image 23354 retrieved from the Centers for Disease Control and Prevention. For the uniform mesh, the edge-size corresponds to 15 pixels. The adaptive was created by controlling the size of the elements based on the difference in the intensity of the pixels.
More Detailed 2D examples of Medical Image Computing can be found here.
Parameters
Note: For the description of the employed sizing function please see the software documentation section of Tessellate3D.
-i, --input [filename] (required)
Input data. It could be an ASCII/Binary NRRD file, an image saved in .vtk format or any other format supported by the ITK library. For more information regarding the file formats supported by the ITK library, see here.
-o, --output [filename] (optional)
The filename of the output mesh saved in VTK format.
(Default: outputMesh.vtk).
-w,--weight-limit [unsigned real] (optional)
Sets the element weight limit for the generated elements that will be used by the sizing function. This parameter limits the difference among the weights within one element. It is designed to give some control of the discretization error with respect to the input data.
(Default: 0.1)
-e,--min-edge [unsigned real] (optional)
Sets the minimum element edge size of the generated elements that will be used by the sizing function. It is designed to be used in conjunction with --weight-limit controlling the size of the generated mesh. Using --min-edge < 1 does not offer significant gain if the pixel size of the input image = 1.
minimum-physical-size of input image = min(spacing * size).
(Default: 1 / 100 * minimum-physical-size)
-u,--uniform (optional)
Creates a uniform mesh instead of an adaptive one. Uses --min-edge value as a constant size constraint.
Computational Nuclear Femtography
A short presentation can be found here.
Tessellate3D
Examples
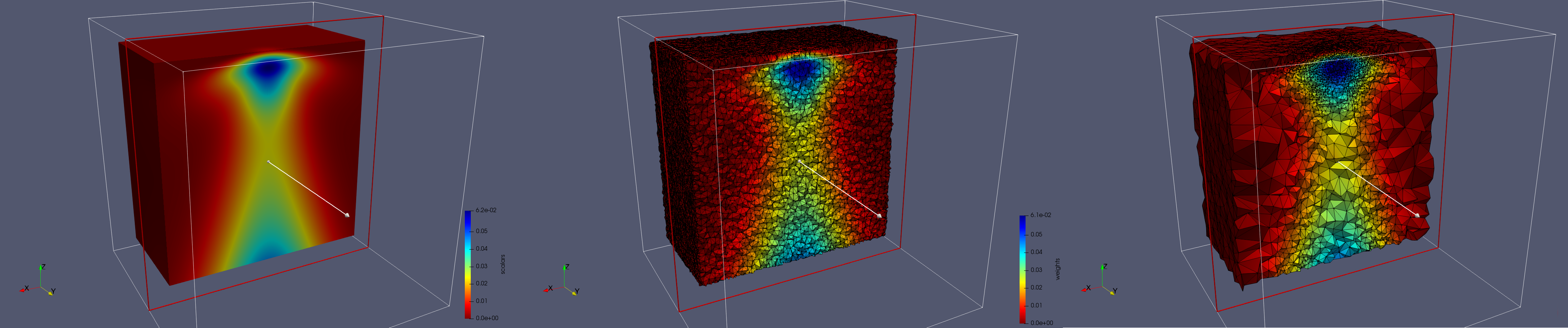
Input image and clipped generated meshes
- Input (left figure): Dimensions (100x100x100) with spacing (1x1x1), Voxels = 1,000,000
- Uniform (middle figure): 768,033 tetrahedra
- Adaptive (right figure): 253,516 tetrahedra
Uniform (middle figure):
docker run -v $(pwd):/data/ crtc_i2m tessellate3d --input ./CNF_Data/3D/CFF_14052019/NT_140519.nrrd --cnf-uniform --output ./NT_140519,d=1,uniform.vtk
Adaptive (right figure):
docker run -v $(pwd):/data/ crtc_i2m tessellate3d --input ./CNF_Data/3D/CFF_14052019/NT_140519.nrrd --cnf-adaptive --weight-limit 0.07 --output ./NT_140519,d=5,wl=0.07,me=1.vtk
More Detailed 3D examples of Computational Nuclear Femtography can be found here.
Parameters
In this domain, some of the parameters of Tessellate3D take on different meanings. For the full range and descriptions of parameters please see the software documentation section of Tessellate3D.
-i, --input [filename] (required)
Input data. It could be an ASCII/Binary NRRD file, an image saved in .vtk format or any other format supported by the ITK library. For more information regarding the file formats supported by the ITK library, see here.
-o, --output [filename] (optional)
The filename of the output mesh saved in VTK format.
(Default: input filename + parameters.vtk).
-t, --threads [unsigned integer] (optional)
Sets the number of threads to be utilized.
(Default: 1).
-d, --delta [unsigned real] (optional)
Controls the size of the elements near the boundary. Smaller values will lead to finer detail close to the boundary (and often to a more accurate boundary representation) but will also lead to a greater mesh size.
minimum-physical-size of input image = min(spacing * size).
(Default (if --cnf-uniform is specified): 1 / 100 * minimum-physical-size).
(Default (if --cnf-adaptive is specified): 1 / 20 * minimum-physical-size).
--cnf-uniform (optional)
Produces uniform size meshes for CNF data. Size of elements is controlled by --delta.
--cnf-adaptive (optional)
Produces adaptive meshes for CNF data. The level of adaptivity is controlled by --weight-limit and --min-edge (see below). For more details about how sizing works see the note at the end of the section.
-w, --weight-limit [unsigned real] (optional)
Sets the element weight limit of the generated elements that will be used by the sizing function. This parameter limits the difference among the weights within one element. It is designed to give some control of the discretization error with respect to the input data.
(Default: 0.1).
-e, --min-edge [unsigned real] (optional)
Sets the minimum element edge size of the generated elements that will be used by the sizing function. It is designed to give some control over the size of the generated mesh. Using --min-edge < 1 does not offer significant gain if the voxel size of the input image = 1.
minimum-physical-size of input image = min(spacing * size).
(Default: 1 / 100 * minimum-physical-size).
-b, --background-value [signed real] (optional)
Sets the voxel value that will be treated as a background value during the image segmentation. If none is desired, enter a value that does not exist in the dataset. In practice, a background value is a value that is ignored by the tessellation procedure. Regions of the tessellation corresponding to the background value will have no elements.
(Default: +oo).
Note: Sizing Function
The input image is used as a background(BG) mesh while refining the mesh. Before the beginning of the refinement, the input image is analyzed, and the global constraints of maximum-element-edge-size and element-weight-range are computed. The refinement algorithm queries the sizing function (SF) to verify whether each element satisfies the global and user constraints. Each time SF is called upon an element, it will check if its size meets the global (maximum-element-edge-size) and user (min-edge) constraints. Consequently, it will create a set SP of sampling points out of the element. This set consists of the element’s vertices, barycenter, and midpoints of vertices and barycenter. The sampling points that are preserved out of SP are those that lie within the BG mesh. Using the BG mesh, a set V of the values of SP is created. Subsequently, the quantity max(V) - min(V) is evaluated. If abs(max(V) - min(V))/element-weight-range exceeds the user constraint weight-limit, the element is split.
Tessellate2D
Examples
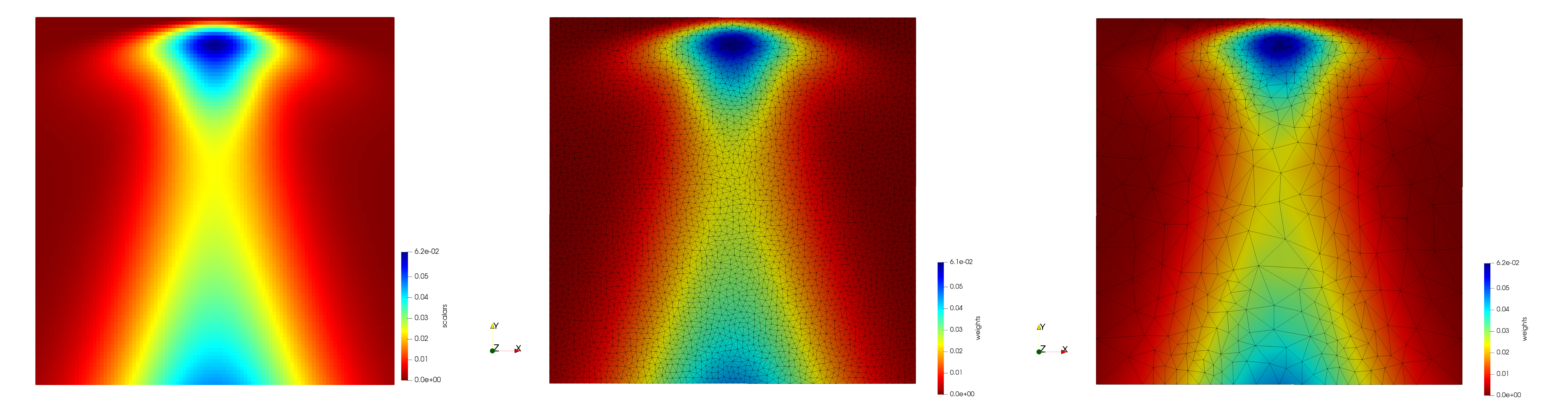
Input image and generated meshes
- Input (left figure) : Dimensions (100x100) with spacing (1x1), Pixels = 10,000
- Uniform (middle figure) : 7,587 triangles
- Adaptive (right figure) : 1,038 triangles
Uniform (middle figure):
docker run -v $(pwd):/data/ crtc_i2m tessellate2d --input ./CNF_Data/2D/NT_140519_50_X.vtk --output NT_140519_X50_me_2_uniform.vtk --min-edge=2 --uniform
Adaptive (right figure):
docker run -v $(pwd):/data/ crtc_i2m tessellate2d --input ./CNF_Data/2D/NT_140519_50_X.vtk --output NT_140519_X50_me_2_wl_1e-1.vtk --min-edge=2 --weight-limit=0.1
More Detailed 2D examples of Computational Nuclear Femtography can be found here.
Parameters
Note: For the description of the employed sizing function please see the software documentation section of Tessellate3D.
-i, --input [filename] (required)
Input data. It could be an ASCII/Binary NRRD file, an image saved in .vtk format or any other format supported by the ITK library. For more information regarding the file formats supported by the ITK library, see here.
-o, --output [filename] (optional)
The filename of the output mesh saved in VTK format.
(Default: outputMesh.vtk).
-w,--weight-limit [unsigned real] (optional)
Sets the element weight limit for the generated elements that will be used by the sizing function. This parameter limits the difference among the weights within one element. It is designed to give some control of the discretization error with respect to the input data.
(Default: 0.1)
-e,--min-edge [unsigned real] (optional)
Sets the minimum element edge size of the generated elements that will be used by the sizing function. It is designed to be used in conjunction with --weight-limit controlling the size of the generated mesh. Using --min-edge < 1 does not offer significant gain if the pixel size of the input image = 1.
minimum-physical-size of input image = min(spacing * size).
(Default: 1 / 100 * minimum-physical-size)
-u,--uniform (optional)
Creates a uniform mesh instead of an adaptive one. Uses --min-edge value as a constant size constraint.
Software Documentation
The CRTC’s image-to-mesh software suite has been packaged into a Docker image for easy distribution and portability across multiple platforms. All that is needed to use it is an OS supporting Docker.
Requirements
- OS: Linux, Windows 10 Pro/Enterprise, MacOS Sierra 10.12+
- Docker
Installing Docker
Official documentation :
Note for Running on Windows
Docker on Windows uses Hyper-V VMs to run Linux containers. By default, the spawned VMs use 2 vCPUs and 2 GB RAM.
If performance is a concern, it is recommended to edit the Docker settings via the GUI to increase the resource allocation for the VMs in order to allow the tessellation3D tool to use more threads.
Getting The Software
The docker image containing the crtc_i2m software suite is located here (restricted access).
Docker Container Instructions
Load the Docker Image.
First of all, the Docker image needs to be loaded. The following command must be used:
docker load --input [DOCKER_IMAGE_TAR]
Note: If the user is not in the
dockergroup, prependingsudoto the above command is necessary.Running
On MacOS/Linux
docker run -v $(pwd):/data/ crtc_i2m <application> [arguments]
On Windows with PowerShell (recommended)
docker run -v ${PWD}:/data/ crtc_i2m <application> [arguments]
Notice that in this case brackets {} are used instead of parenthesis ()
On Windows with the command line (cmd)
docker run -v %cd%:/data/ crtc_i2m <application> [arguments]
Where <application> is one of the currently available tools : tessellate3d, tessellate2d, convert_image or partonsToNRRD.
Tessellate3D (PODM 3D)
The main component of the software suite is a 3D tessellation software called PODM3D, which is a parallel Image-to-Mesh conversion algorithm with quality and fidelity guarantees and is capable of generating unstructured tetrahedral meshes out of 3D structured data.
The output meshes are in the VTK format and can be visualized using the open-source software Paraview.
A quick way to view all available parameters and brief descriptions for them is to pass the -h, --help flag to tessellate3d.
Below is the detailed information about the parameters of tessellate3d:
I/O Parameters
-i, --input [filename] (required)
Input data. It could be an ASCII/Binary NRRD file, an image saved in .vtk format or any other format supported by the ITK library. For more information regarding the file formats supported by the ITK library, see here.
-o, --output [filename] (optional)
The filename of the output mesh saved in VTK format.
(Default: input filename + parameters.vtk).
-c, --plc [filename] (optional)
If given, the surface of the produced mesh will be saved into filename in the VTK format.
-r, --file-format {0,1} (optional)
Sets file format of outputFiles. 0 for ASCII, and 1 for BINARY.
(Default: ASCII).
Performance Parameters
-t, --threads [unsigned integer] (optional)
Sets the number of threads to be utilized. The upper bound of the number of threads that should be utilized is equal to the number of cores that you have. If the number of threads is greater than 1, the produced meshes may differ in terms of the number of elements up to 3% due to the nature of the employed parallelism technique.
(Default: 1).
-p, --thread-pinning {0, 1} (optional)
Controls the pinning of threads to cores. 0 doesn’t pin threads to cores (experimental), and 1 pins threads to cores.
(Default: 1).
-m, --memory-limit [unsinged integer in MB] (optional)
Constrains the amount of memory in MB that will be used.
(Default: 70% of the free memory).
Geometry Parameters
-d, --delta [unsigned real] (optional)
Controls the density of surface approximation. Smaller values will lead to denser approximation close to the surface (and often to a more accurate surface representation), but will also lead to greater mesh size. The same delta value is used for every tissue of interest. A smaller delta value should be used if at least one of the tissues of interest is not recovered after the Meshing Procedure. For uniform meshes (i.e. no --volume-grading), the total number of elements is related to delta through an inverse cubic law (approximately):
The maximum suggested value for delta is 1 / 5 of the minimum-physical-size of the input image, where minimum-physical-size = min(spacing * size).
(Default (if --sizing-function is not specified): 1 / 100 * minimum-physical-size).
(Default (if --sizing-function is specified): 1 / 20 * minimum-physical-size).
Delta is an essential parameter that is involved in every step of the PODM algorithm and guarantees the quality and fidelity of the produced mesh. If using d_1 creates N_1 elements then d_1/2 will create (approximately) 8 = 2^3 times more elements while 2 * d_1 will generate 1/8N_1 elements. A table which shows the relation between delta and mesh size can be found in the Appendix. For more information on how delta affects the density of the sampling, see Guaranteed Quality Tetrahedral Delaunay Meshing for Medical Images.
-g, --volume-grading (optional)
Enables the grading of the volume of the mesh. By default, the value of delta controls both the surface approximation and the size of the elements. Using this flag the value of delta will control only the surface approximation resulting in elements of higher volume inside the domain.
Pre-processing Parameters
-s, --image-segmentation (optional)
Performs Image Segmentation on non-segmented/unlabeled images using a given background value. It should not be used for input images that are already segmented/labeled. Uses --background-value as an optional parameter.
-b, --background-value [signed real] (optional)
Sets the voxel value that will be treated as a background value during the image segmentation. If none is desired, enter a value that does not exist in the dataset. In practice, a background value is a value that is ignored by the tessellation procedure. Regions of the tessellation corresponding to the background value will have no elements.
(Default: +oo).
--include-labels [unsigned char list] (optional)
Labels that will be included in the mesh. If not specified, all labels will be included in the mesh. Cannot be used together with --exclude-labels.
--exclude-labels [unsigned char list] (optional)
Labels that will be excluded from the mesh. Keeps all labels in the mesh except the ones specified. If not specified, all labels will be included in the mesh. Cannot be used together with --include-labels.
Mesh Sizing Parameters
-f, --sizing-function (optional)
Enables the sizing-function that is described below. Uses --weight-limit and --min-edge as optional parameters.
--sizing-function needs to be used in conjunction with --volume-grading so that the volume’s refinement will be mainly controlled by the sizing-function.
The input image is used as a background(BG) mesh while refining the mesh. Before the beginning of the refinement, the input image is analyzed, and the global constraints of maximum-element-edge-size and element-weight-range are computed. The refinement algorithm queries the sizing function (SF) to verify whether each element satisfies the global and user constraints. Each time SF is called upon an element, it will check if its size meets the global (maximum-element-edge-size) and user (min-edge) constraints. Consequently, it will create a set SP of sampling points out of the element. This set consists of the element’s vertices, barycenter, and midpoints of vertices and barycenter. The sampling points that are preserved out of SP are those that lie within the BG mesh. Using the BG mesh, a set V of the values of SP is created. Subsequently, the quantity max(V) - min(V) is evaluated. If abs(max(V) - min(V))/element-weight-range exceeds the user constraint weight-limit, the element is split.
The described sizing function was designed for the Computational Nuclear Femtography data. Nonetheless, another sizing function could be designed and employed if requested.
-w, --weight-limit [unsigned real] (optional)
Sets the element weight limit of the generated elements that will be used by the sizing function. This parameter limits the difference among the weights within one element. It is designed to give some control of the discretization error with respect to the input data.
(Default: 0.1).
-e, --min-edge [unsigned real] (optional)
Sets the minimum element edge size of the generated elements that will be used by the sizing function. It is designed to give some control over the size of the generated mesh. Using --min-edge < 1 does not offer significant gain if the voxel size of the input image = 1.
minimum-physical-size of input image = min(spacing * size).
(Default: 1 / 100 * minimum-physical-size).
Post-processing Parameters
-l, --linear-interpolation (optional)
Performs Linear Interpolation over the points of the produced mesh using the input image. It should not be used for input images that are already segmented/labeled.
Statistics Parameters
--thread-statistics (optional)
Computes and prints thread statistics.
--mesh-statistics (optional)
Computes and prints mesh statistics.
--all-statistics (optional)
Computes and prints all statistics (thread and mesh).
Special Parameters
--cnf-uniform (optional)
Activates the flags --image-segmentation, and --linear-interpolation which are required for producing uniform size meshes for CNF data. Size of elements is controlled by --delta.
--cnf-adaptive (optional)
Activates the flags --image-segmentation, --volume-grading, --sizing-function, and --linear-interpolation which are required for producing adaptive meshes for CNF data. The level of adaptivity is controlled by --weight-limit and --min-edge.
Miscellaneous Parameters
-v, --verbosity-level {0, 1, 2} (optional)
Controls the level of output text verbosity. 0 produces no output, 1 produces standard output, and 2 produces extensive output.
(Default: 1).
--version (optional)
Show PODM3D version and exit.
Tessellate2D (Triangle)
Tessellate2D is a modified version of the Triangle software for 2D tessellation.
The output meshes are in the VTK format and can be visualized using the open-source software Paraview.
Note: For the description of the employed sizing function please see the software documentation section of Tessellate3D.
Below is the detailed information about the parameters of tessellate2d:
-i, --input [filename] (required)
Input data. It could be an ASCII/Binary NRRD file, an image saved in .vtk format or any other format supported by the ITK library. For more information regarding the file formats supported by the ITK library, see here.
-o, --output [filename] (optional)
The filename of the output mesh saved in VTK format.
(Default: outputMesh.vtk).
-r, --file-format {0,1} (optional)
Sets file format of outputFiles. 0 for ASCII, and 1 for BINARY.
(Default: ASCII).
-w,--weight-limit [unsigned real] (optional)
Sets the element weight limit for the generated elements that will be used by the sizing function. This parameter limits the difference among the weights within one element. It is designed to give some control of the discretization error with respect to the input data.
(Default: 0.1)
-e,--min-edge [unsigned real] (optional)
Sets the minimum element edge size of the generated elements that will be used by the sizing function. It is designed to be used in conjunction with --weight-limit controlling the size of the generated mesh. Using --min-edge < 1 does not offer significant gain if the pixel size of the input image = 1.
minimum-physical-size of input image = min(spacing * size).
(Default: 1 / 100 * minimum-physical-size)
-u,--uniform (optional)
Creates a uniform mesh instead of an adaptive one. Uses --min-edge value as a constant size constraint.
-v,--verbose-level [0,1] (optional)
Controls the level of output text verbosity. 0 produces no output, and 1 produces standard output.
(Default: 1).
Convert Image
convert_image serves as a utility to convert input data between different types of image formats. It allows converting the input data between different image types supported by ITK enabling in some cases more post-processing filters in Paraview (e.g. contour plots). However, converting them using the following command enables all relevant image filters in Paraview.
docker run -v $(pwd):/data/ crtc_i2m convert_image input_image.nrrd output_image.vtk
Partons To NRRD
partonsToNRRD serves as a utility to create images out of partons data
docker run -v $(pwd):/data/ crtc_i2m partonsToNRRD partonsFile.dat columnId xBins yBins zBins
Appendix
How mesh size relates to delta
Input Image: NT_140519.nrrd
- The size of the input image is: (100x100x100)
- Spacing of the input image: (1x1x1)
- MinimumPhysicalSize = 100 * 1 = 100
- Maximum suggested delta value is: 100 / 5 = 20
| Delta | # Vertices | # Tetrahedra |
|---|---|---|
| 20 | 147 | 430 |
| 10 | 563 | 1,780 |
| 5 | 2,686 | 10,210 |
| 2.5 | 13,366 | 56,274 |
| 1.25 | 79,086 | 401,610 |
| 0.625 | 520,538 | 2,922,427 |