Difference between revisions of "Medical Imaging Example Meshes"
From crtc.cs.odu.edu
Spyridon97 (talk | contribs) (→2D Example Meshes) |
Spyridon97 (talk | contribs) (→2D Example Meshes) |
||
| Line 2: | Line 2: | ||
=2D Example Meshes= | =2D Example Meshes= | ||
The directory containing the 2D input data is located in the 2D folder of [https://odu.box.com/s/wd0i18giti2zo330y19xhkqu8wjzgf7j Medical_Imaging_Data]. | The directory containing the 2D input data is located in the 2D folder of [https://odu.box.com/s/wd0i18giti2zo330y19xhkqu8wjzgf7j Medical_Imaging_Data]. | ||
| + | |||
The image data were acquired from [https://www.cdc.gov/media/subtopic/images.htm Centers for Disease Control and Prevention] | The image data were acquired from [https://www.cdc.gov/media/subtopic/images.htm Centers for Disease Control and Prevention] | ||
Revision as of 08:33, 14 March 2020
Contents
2D Example Meshes
The directory containing the 2D input data is located in the 2D folder of Medical_Imaging_Data.
The image data were acquired from Centers for Disease Control and Prevention
3D Example Meshes
The directory containing the 3D input data is located in the 3D folder of Medical_Imaging_Data.
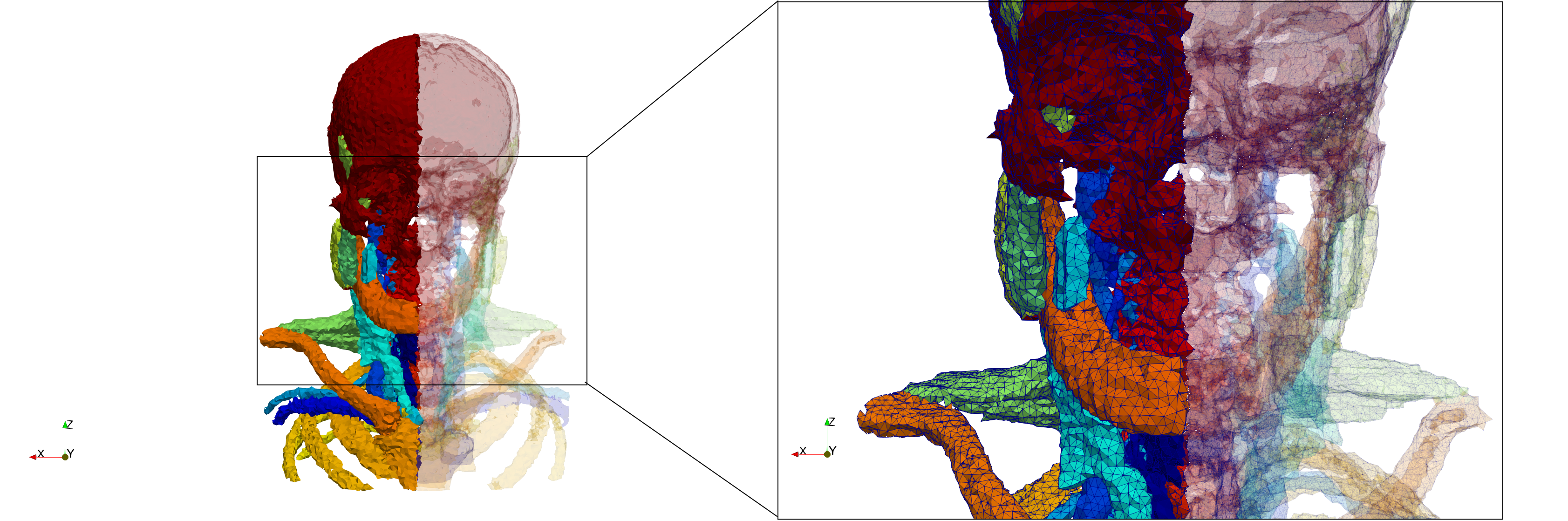
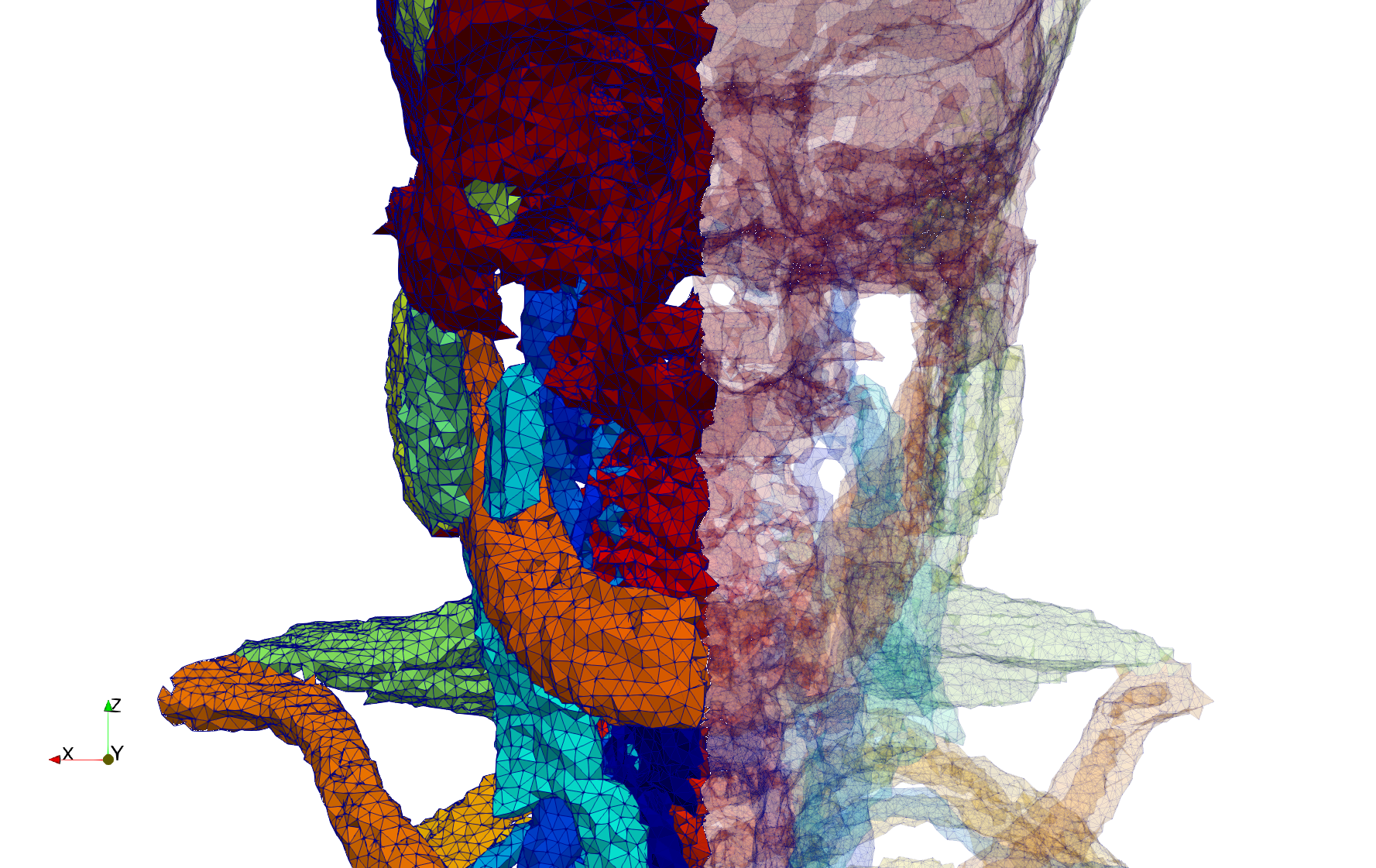
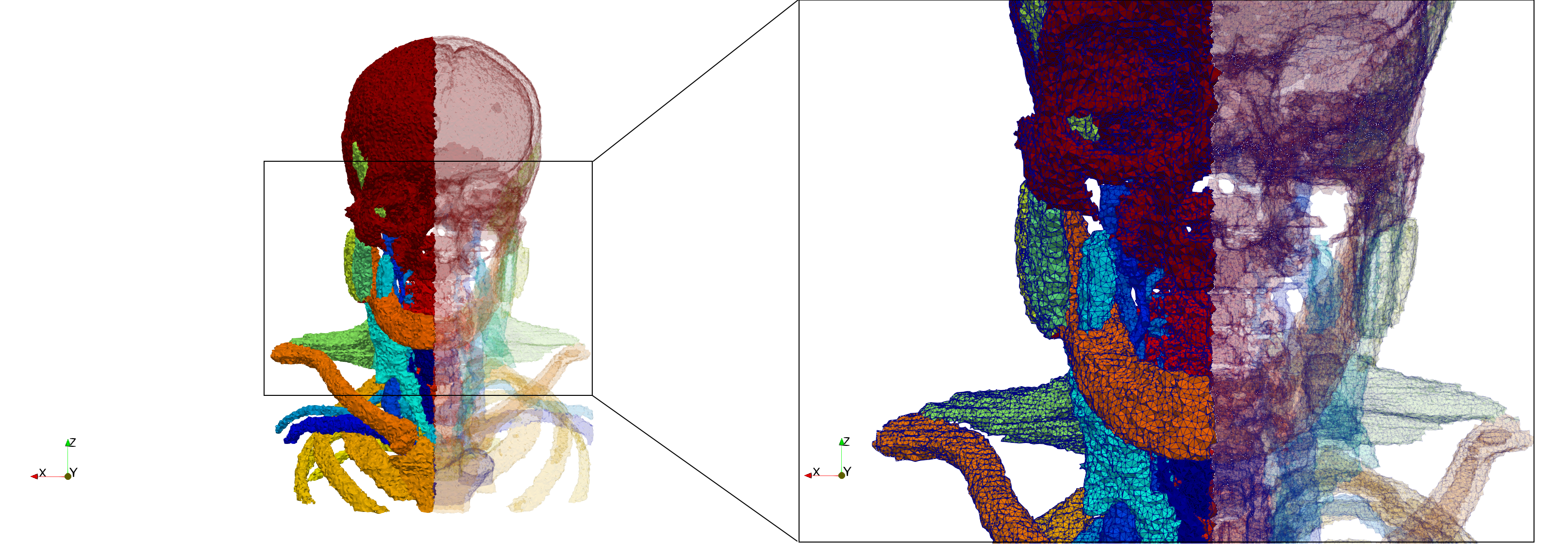
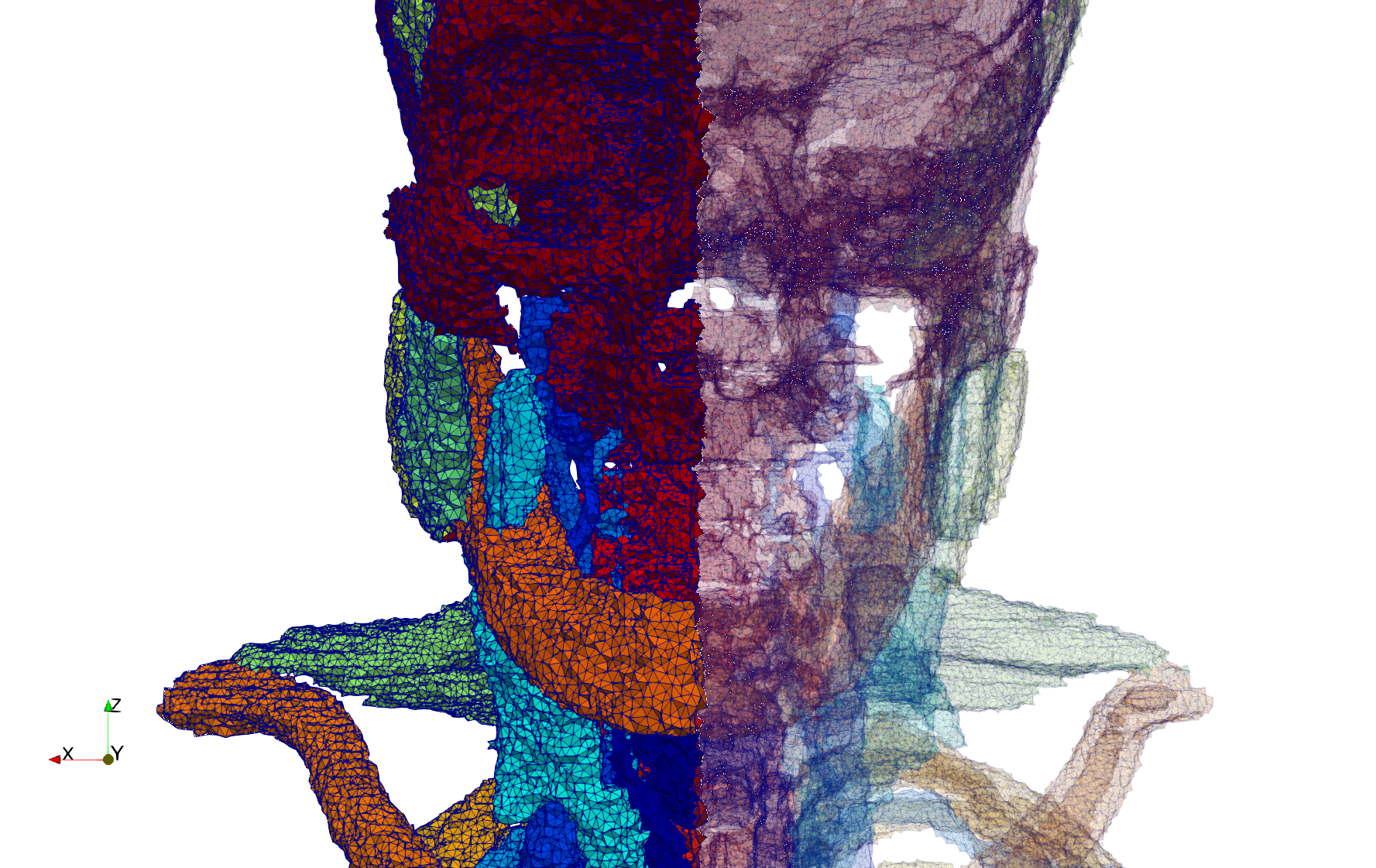
Head-Neck
- Input image : Dimensions (255x255x229) with spacing (0.976562x0.976562x1.40002)
- Uniform with Delta = default: 205,416 tetrahedra
- Uniform with Delta = 1.5: 770,853 tetrahedra
Commands to generate meshes :
Uniform with Delta = default : Output Mesh
docker run -v $(pwd):/data/ crtc_i2m tessellate3d --input ./Medical_Imaging_Data/3D/Head-Neck.mha --output ./Head-Neck,d=2.49023.vtk
Uniform with Delta = 1.5 : Output Mesh
docker run -v $(pwd):/data/ crtc_i2m tessellate3d --input ./Medical_Imaging_Data/3D/Head-Neck.mha --delta 1.5 --output ./Head-Neck,d=1.5.vtk
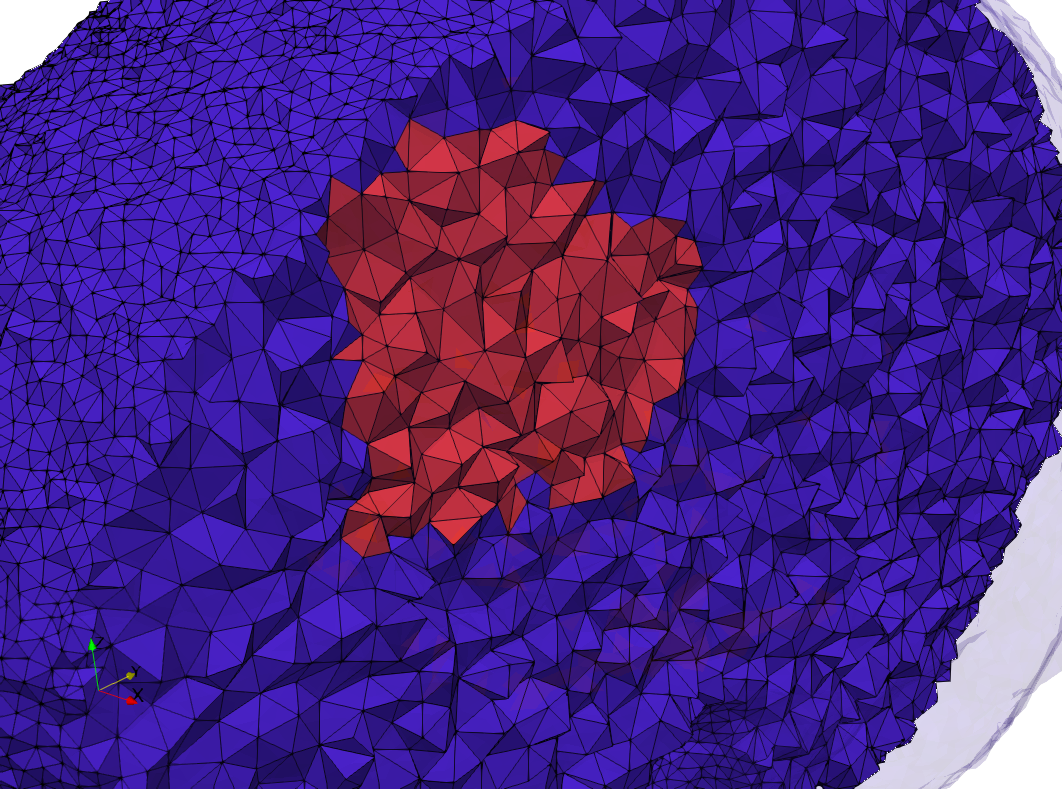
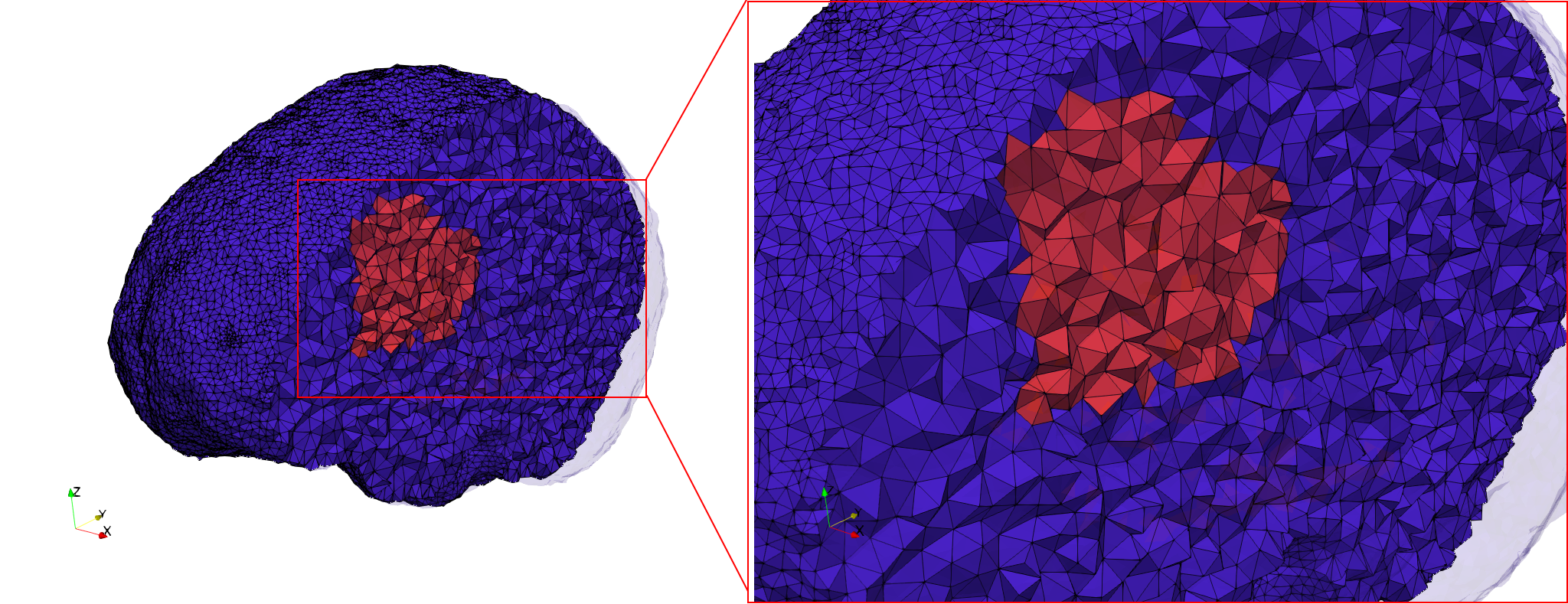
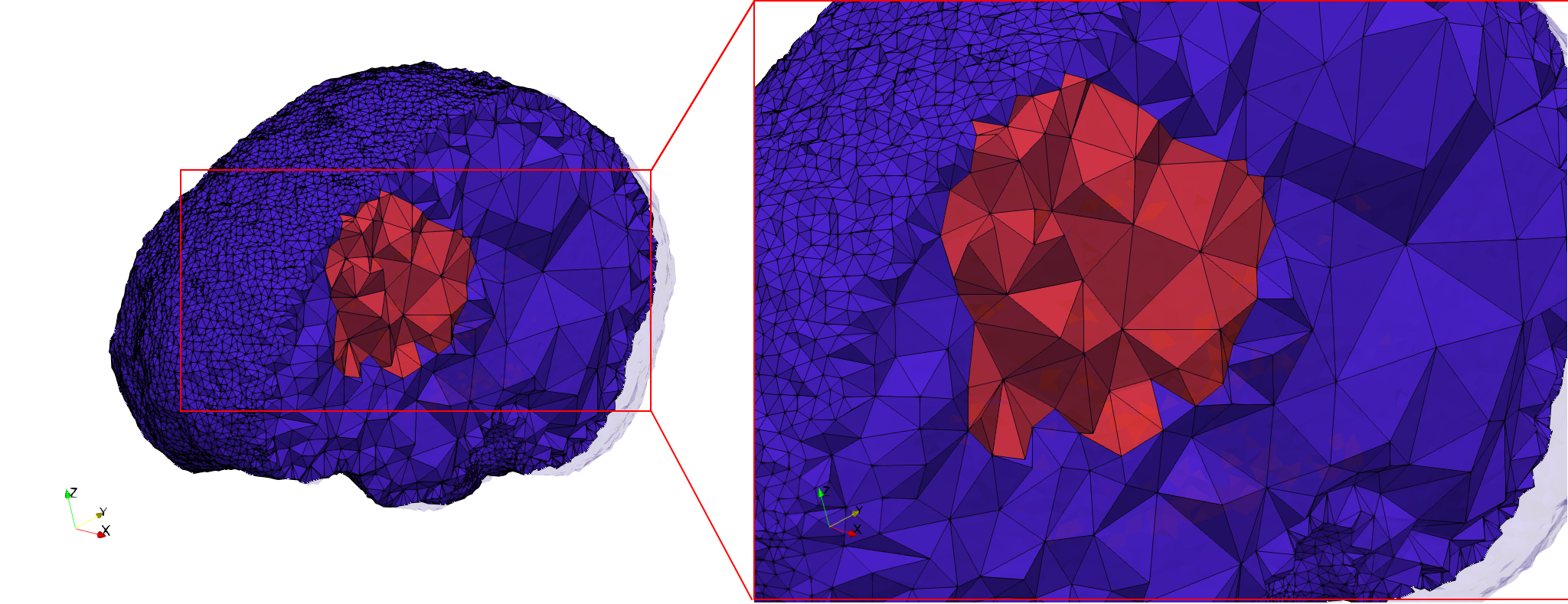
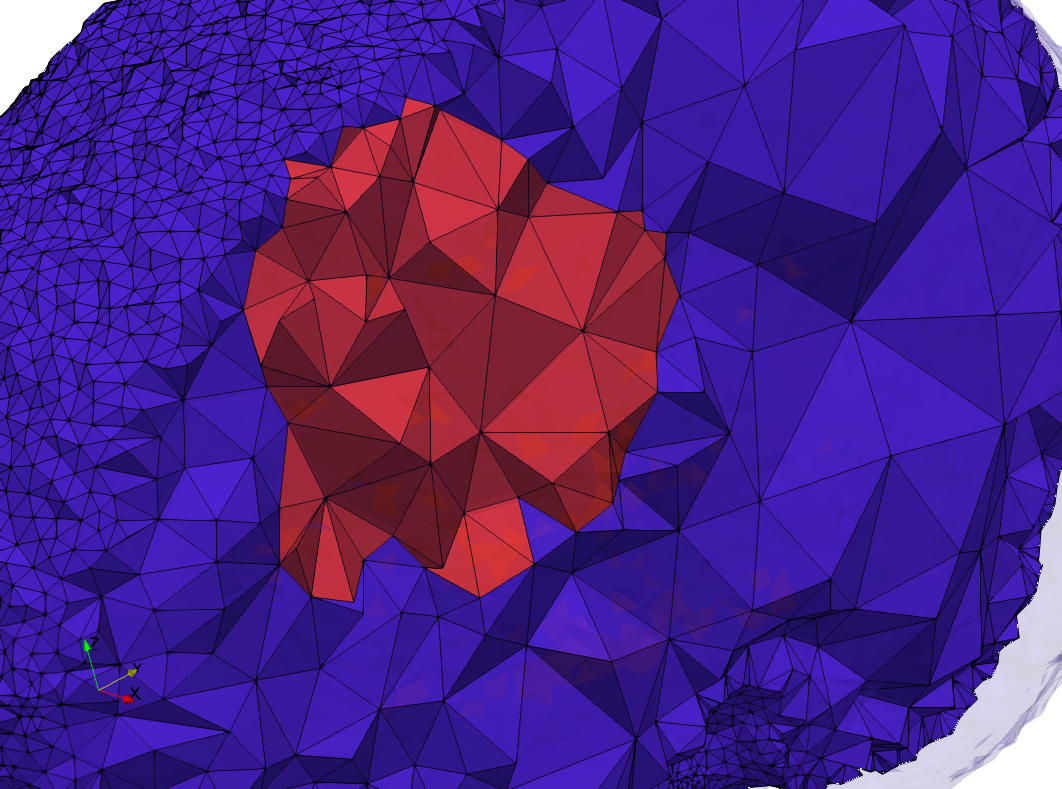
Brain-With-Tumor-Case17
- Input image : Dimensions (448x512x176) with spacing (0.488281x0.488281x1)
- Uniform with Delta = default: 222,540 tetrahedra
- Graded with Delta = default: 94,383 tetrahedra
Commands to generate meshes :
Uniform with Delta = default : Output Mesh
docker run -v $(pwd):/data/ crtc_i2m tessellate3d --input ./Medical_Imaging_Data/3D/Brain-With-Tumor-Case17.nii --output ./Brain-With-Tumor-Case17,d=1.76001.vtk
Graded with Delta = default : Output Mesh
docker run -v $(pwd):/data/ crtc_i2m tessellate3d --input ./Medical_Imaging_Data/3D/Brain-With-Tumor-Case17.nii --volume-grading --output ./Brain-With-Tumor-Case17,d=1.76001,graded.vtk
Knee-Char
- Input image : Dimensions (512x512x119) with spacing (0.27734x0.27734x1)
- Uniform with Delta = default case: 386,869 tetrahedra
- Graded with Delta = default case: 274,309 tetrahedra
Commands to generate meshes :
Uniform with Delta = default : Output Mesh
docker run -v $(pwd):/data/ crtc_i2m tessellate3d --input ./Medical_Imaging_Data/3D/Head-Neck.mha --output ./Knee-Char,d=1.19.vtk
Graded with Delta = default : Output Mesh
docker run -v $(pwd):/data/ crtc_i2m tessellate3d --input ./Medical_Imaging_Data/3D/Head-Neck.mha --volume-grading --output ./Knee-Char,d=1.19,graded.vtk
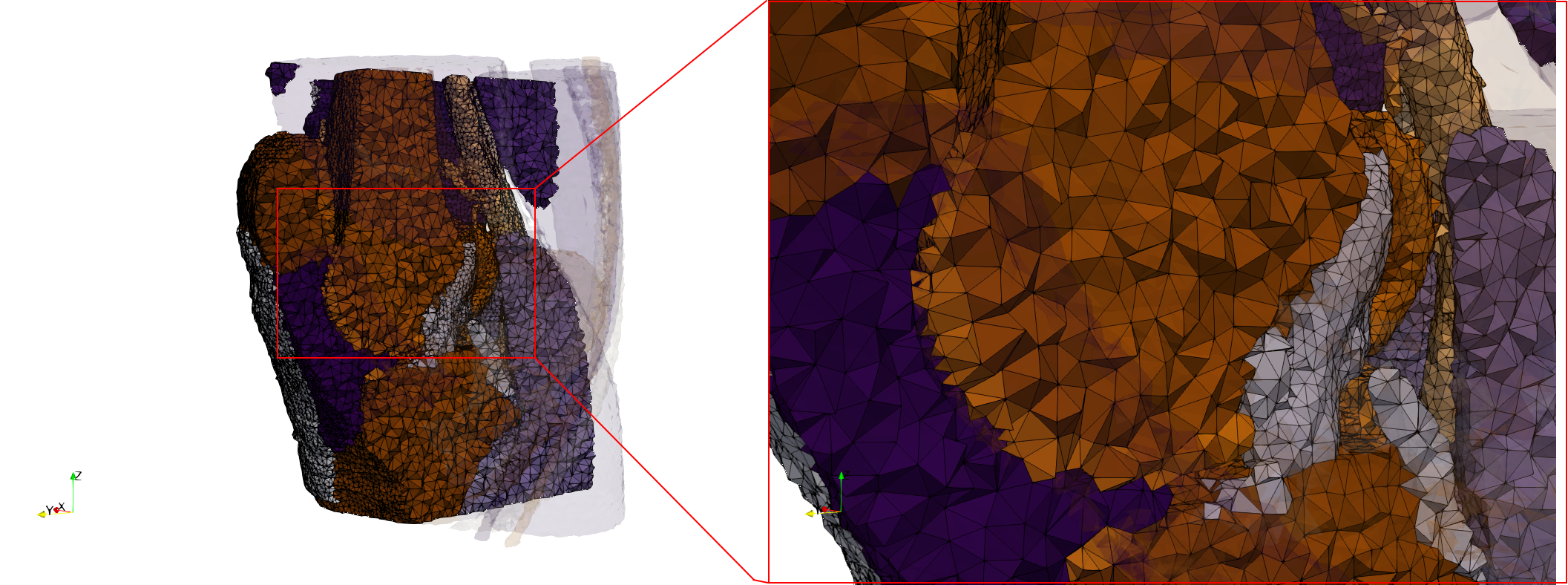
Ircad2
- Input image : Dimensions (512x512x219) with spacing (0.976562x0.976562x1.40002)
- Uniform with Delta = 2 case: 5,031,442 tetrahedra
- Graded with Delta = 1 case: 6,072,751 tetrahedra
Commands to generate meshes :
Uniform with Delta = 2 : Output Mesh
docker run -v $(pwd):/data/ crtc_i2m tessellate3d --input ./Medical_Imaging_Data/3D/Ircad2.nrrd --delta 2 --output ./Ircad2,d=2.vtk
Graded with Delta = 1 : Output Mesh
docker run -v $(pwd):/data/ crtc_i2m tessellate3d --input ./Medical_Imaging_Data/3D/Ircad2.nrrd --delta 1 --volume-grading --output ./Ircad2,d=1,graded.vtk